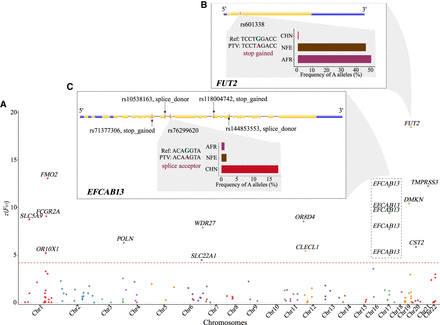
PTVs with high population differentiation in allelic frequency between Chinese (CHN), non-Finnish European (NFE), and African (AFR) populations. (A) Target-wide Z(FST) Manhattan plot. Note that Weir and Cockerham FST was normalized and transformed into Z values and the Bonferroni correction set the significant Z(FST) cutoff at 4.3. (B) (Top) Gene structure of FUT2. (Bottom) Nucleotide sequence and allele frequency distribution of a stop-gained G-A PTV (rs601338) with a Z(FST) of 18.5. (C) (Top) Gene structure of EFCAB13 with five PTVs of significant Z(FST). (Bottom) Nucleotide sequence and allele frequency distribution of a splice-acceptor G-A PTV (rs76299620) with a Z(FST) of 12.2.











