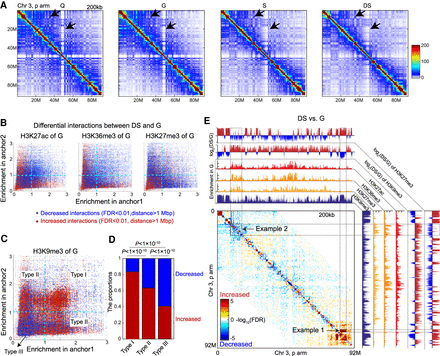
H3K9me3-associated heterochromatin of G gains long-range interactions. (A) Contact maps of the p arm of Chr 3 across four cell states. The arrows on the plot point out positions of some differences. (B) Scatterplots of histone modification (H3K27ac, H3K36me3, and H3K27me3) enrichments of G for two genomic anchors of differential interactions. Each point represents a significantly differential interaction (FDR < 0.01, distance > 1 Mbp). Red represents increased interactions in DS, and blue denotes decreased interactions. Each interaction has two genomic anchors, whose histone modification enrichments of G are shown as x and y coordinates, respectively. These plots could inform the relationship between the enrichment of histone modifications in two genomic anchors and the directions of differential interactions. (C) Scatterplots of H3K9me3 occupation of G in anchors of differential interactions. Interactions are classified into three types (Type I, Type II, and Type III). Type I represents interactions of two H3K9me3-enriched genomic regions; Type II represents interactions between H3K9me3-enriched regions and H3K9me3-depleted regions; Type III represents interactions of two H3K9me3-depleted regions. (D) The composition of significantly increased and decreased interactions for three interaction types. P-values were calculated by a χ2 test. (E) Differential interactions of the p arm of Chr 3. Examples 1 and 2 show increased interactions between heterochromatin and decreased interactions between euchromatin, respectively. The tracks of histone modifications of G and fold changes of H3K9me3 and H3K27me3 between DS and G are shown. Although Example 1 resides in pericentromeric regions, increased heterochromatin interactions also occupy other genomic regions.











