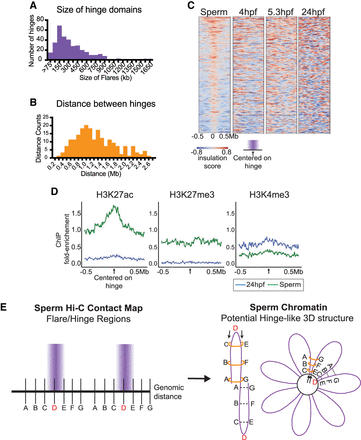
Global characterization of “hinge-like” domains in sperm chromatin architecture. (A) Histogram to depict the width distribution of the 333 hinges. (B) Histogram to depict the average distances between adjacent hinges. (C) Insulation scores at hinge-like domains in sperm, and insulation scores at the corresponding regions in embryos at 4, 5.3, and 24 hpf. Positive insulation (red) indicates increased contacts, and negative insulation (blue) indicates a lack of contacts. (D) Metaplot of log2 fold enrichment of histone H3K27ac (Zhang et al. 2018), H3K27me3 (Irimia et al. 2012), and H3K4me3 (Zhang et al. 2016) ChIP-seq signal over input for sperm (green) and 24 hpf (blue) centered on the hinge-like domains. (E) Two schematics depict two speculative models of zebrafish sperm chromatin and hinge architecture. In both models, the sperm DNA (which is nucleosome-packaged, not protamine-packaged) is arranged into arrays of consecutive loops/petals, possibly similar to the loops described for condensed mitotic chromosomes in somatic cells (Gibcus et al. 2018). Within each petal DNA positions A–G represent the repeated “hinge” unit within each petal, with “D” representing the hinge center, and the segments A–C and E–G representing the edges of the hinge “petals.” A key feature of the contact map data is that locations equidistant from position D show increased interaction. Two models are presented to achieve this: (1) interactions are caused by contacts within each petal (left), or (2) interactions are caused by contacts between two petals (right). In both models, we propose a constraint on topology, which might involve the loading of ring-like proteins (e.g., condensin or cohesin complexes) at the hinge position D (left) which, together with the fixed hinge position D, create the hinge-like domain through loop extrusion. Ring-like proteins are represented by orange rings. Arrows on the left indicate potential locations where ring-like proteins might load, at hinge position D, to help form stable hinges.











