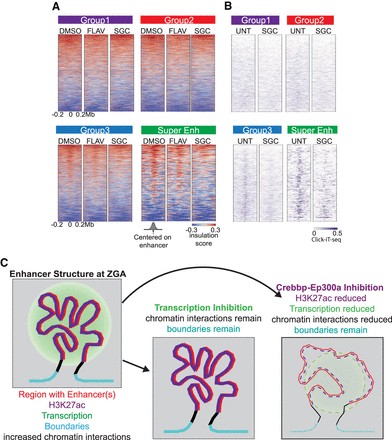
Inhibition of Crebbp/Ep300a causes loss of chromatin architecture around the established super-enhancers at 4 hpf. (A) Heatmaps of insulation score for drug-treated embryos. Treatments involve DMSO (vehicle), flavopiridol (FLAV), SGC-CP30 (SGC) for 4 h (which causes a developmental arrest for FLAV and SGC). Respective enhancer groups are ranked as in Figure 3B. Positive insulation (red) indicates increased contacts, and negative insulation (blue) indicates a lack of contacts. (B) Click-iT-seq of 4 hpf untreated (UNT) and SGC-CP30 (SGC, purple) (Chan et al. 2019) heatmaps centered on enhancers of each respective enhancer group ranked as in Figure 4A. (C) Model depicting the features present at regions displaying structure/positive interaction. Features displayed include enhancers (red), elevated histone H3K27ac (purple), active transcription (green circle), defined boundaries (cyan), and increased chromatin interactions as detected by positive interaction scores in Hi-C contact maps at both 4 and 5.3 hpf. Regions with increased interactions are typically coated with histone H3K27ac. These interactions and boundaries persist upon inhibition of RNA Pol II initiation at 4 hpf. In contrast, these contacts between boundaries are lost upon inhibition of Crebbp/Ep300a (lowering histone H3K27ac [dashed]) leading to decreased transcription and loss of higher-order chromatin structure; however, boundaries remain stable.











