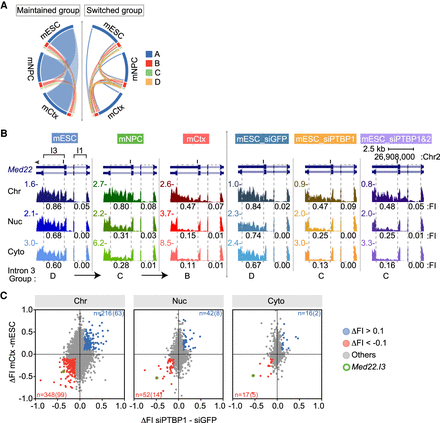
Regulation of intron retention and chromatin association during neuronal development. (A) Circos plot (Krzywinski et al. 2009; Gu et al. 2014) of intron group changes between cell types (mESCs, mNPCs, and mCtx neurons). Introns not changing groups are on the left. Introns switching groups between cell types on the right. (B) Genome browser tracks of Med22 during neuronal differentiation (left three panels) and after Ptbp knockdown in mESCs (right three panels). Dashed boxes indicate U introns with measured FI values (introns 1 and 3) under each track. Group classification of intron 3 is at the bottom. (C) Scatter plots of FI change between mESCs and neurons (mCtx) plotted for each fraction against FI change after Ptbp1 knockdown in mESCs. Introns with Δ FI < −0.1 in both conditions are in red and with Δ FI > 0.1 in blue. The number of introns showing these changes with the number carrying PTBP1 iCLIP tags in parentheses, is above and below (Linares et al. 2015). Intron 3 of Med22 is circled in green.











