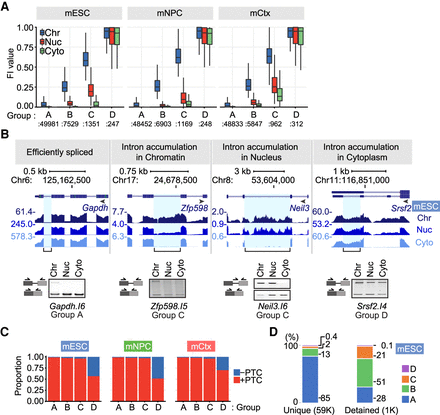
Intron groups defined by their retention level and fractionation behavior. (A) X-means clustering was applied to intron FI values and fraction enrichment in mESCs, mNPCs, and mCtx neurons. The FI distribution for introns in each subcellular fraction and group is shown. (B) Genome browser tracks (top) and RT-PCR validation (bottom) of representative transcripts in mESCs. Validated introns are indicated by a blue highlight and a bracket below. Gel images are one of three biological replicates. (C) The proportion of introns containing a PTC in frame with the upstream sequence is shown for each cluster and cell type. (D) Percentage of introns in each group for U introns from mESCs and for detained introns within the U intron set (Boutz et al. 2015).











