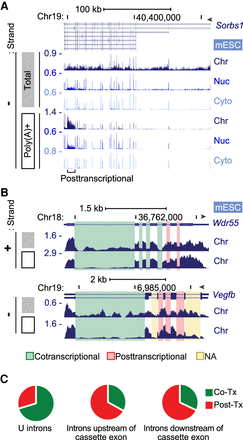
Cotranscriptional and posttranscriptional intron excision. (A) Genome browser tracks of the Sorbs1 locus in mESCs. Total chromatin RNA (gray box) shows intron reads, but the poly(A)+ RNA (open box) shows primarily exon reads except one posttranscriptional intron. (B) Genome browser tracks of chromatin RNA at the Wdr55 and Vegfb loci in mESCs. Total (gray box) and poly(A)+ (open box) are shown, with cotranscriptionally and posttranscriptionally spliced introns highlighted in green and red, respectively. Yellow highlighted introns were not analyzable owing to multiple processing patterns. (C) Proportions of co- and posttranscriptional splicing for 49,692 U introns in mESCs, using criteria described in Supplemental Figure S4, C through E. Introns upstream of (2779) and downstream from (2744) simple cassette exons were similarly analyzed.











