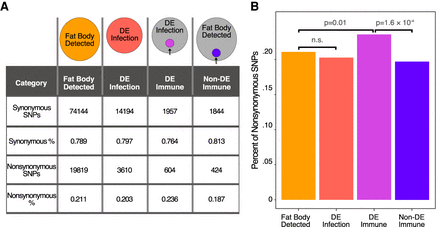
There is a greater proportion of nonsynonymous SNPs in previously identified immune-responsive genes. (A) To look for the prevalence of nonsynonymous SNPs in our genotypes and genes of interest, we defined four gene sets. Among genes detected in the fat body samples, we separated genes into those that were differentially expressed in response to either infection (DE infection) and those that were not (fat body detected). Within the fat body–detected genes, we defined previously identified immune genes showing no differential expression in response to infection (non-DE immune), and among the DE infection genes, we refined the gene list to include previously identified immune genes (DE immune). The numbers indicate the total number of SNPs found in each gene set and the percentages of synonymous and nonsynonymous SNPs. (B) DE immune genes have a higher proportion of nonsynonymous SNPs than the fat body–expressed genes, which suggests they may carry function-altering SNPs at a higher rate than the fat body–expressed genes. P-values are Bonferroni-corrected from chi-square tests with the proportion of nonsynonymous SNPS relative to the fat body–expressed gene set.











