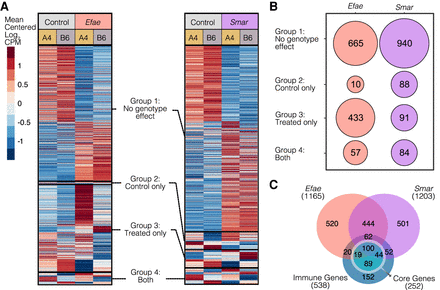
The A4 and B6 D. melanogaster lines have variation in their response to Gram-positive E. faecalis (Efae) infection and Gram-negative S. marcescens (Smar). (A) We measured expression in the fat bodies of the A4 and B6 lines infected with Efae or with Smar, 3 h post infection. We found 1165 and 1205 differentially expressed genes in response to infection to Efae and Smar, respectively, relative to control samples. Mean centered log2 average CPM values for each condition are displayed. We categorized the infection-responsive genes into four groups, based on their differential expression between the two fly genotypes: genes showing no genotype-specific expression (Group 1), genes showing genotype-specific expression only in the control condition (Group 2), genes showing genotype-specific expression only in the infected condition (Group 3), and genes showing genotype-specific expression in both the control and infected conditions (Group 4). (B) Among genes showing genotype effects, the majority of genes in Efae fell into the Group 2 classification, indicating a large amount of genotype-specific expression variation is revealed upon infection with Efae. Among Smar-responsive genes, roughly equal numbers show expression differences between the genotypes before (Group 2), after (Group 3), and both before and after infection (Group 4). (C) We intersected the genes we identified as differentially expressed in response to infection and a list of previously published immune-responsive genes. More than half of the verified immune genes were identified as differentially expressed in the abdominal fat body, with half of these immune genes being shared between conditions. Among these previously identified immune genes, core genes are differentially expressed across all infections. We detected ∼40% of the core set as differentially expressed in both our infection conditions, despite differences in the genetic background, tissue type, and time point used in our study versus previous work.











