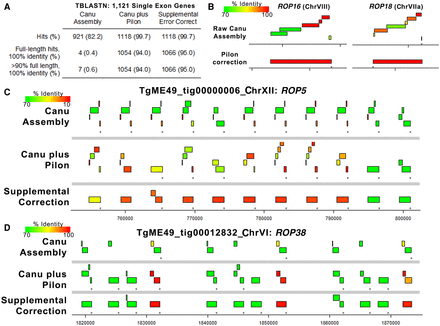
Error polishing with Pilon is effective for most single-copy genes, but resolution of tandem gene expansion errors requires supplemental correction. (A) We identified 1121 single-exon genes with predicted proteins that mapped with 100% identity and 100% coverage using TBLASTN against the ToxoDB-48 ME49 genome. Then, we mapped these against the raw assembly generated by Canu, the polished assembly generated by four rounds of Pilon, and after four rounds of supplemental error correction. Pilon error correction was sufficient for perfect mapping of 94% of the query single-exon genes (compared with only 0.4% for the raw Canu assembly), and supplemental error correction only increased this mapping percentage slightly. (B) Plots representing TBLASTN analysis of protein sequences from two single-copy genes showing the improved mapping achieved by Pilon-based error correction. Mapping identity is indicated by the color of the box representing the alignment. (C,D) Plots representing protein-coding sequences from the ROP5 (C) or ROP38 (D) gene mapped using TBLASTN against the raw Canu-only assembly, the Pilon-corrected assembly, and the region corrected using our supplemental approach tailored to tandem gene arrays. Both loci have multiple pseudogenes in the Canu-only and Canu-plus Pilon assemblies, but many of these errors are removed upon supplemental correction. The presence of a pseudogene in the ME49 ROP5 locus has been predicted before based on direct sequencing, suggesting that this may represent the most accurate version of the ME49 ROP5 locus sequenced to date.











