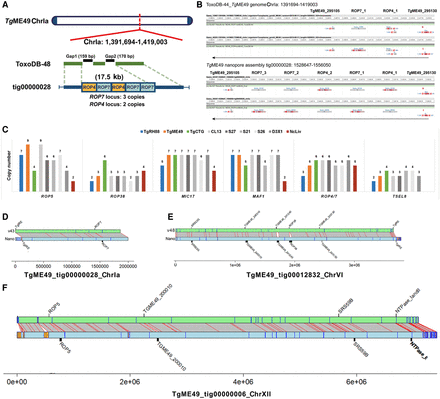
Long-read assembly resolves duplicated locus structure in T. gondii genome. (A) Two unresolved scaffold gaps on Chr Ia in the ToxoDB-48_TgME49 genome span a 17.5-kb tandem repeat containing multiple copies of ROP4 and ROP7. The ROP4/7 gaps are closed by the TgME49 long-read assembly (TgME49_tig00000028), revealing a tandem array of five copies of this gene in the order shown. (B) BLASTN alignment of the ROP4/ROP7 coding sequence in the ToxoDB-48_TgME49 genome (upper panel) and the TgME49 long-read assembly (lower panel). (C) Copy number determination at six canonical tandem gene arrays across eight T. gondii strains and one N. caninum strain. Data from CL13, S27, S21, and S26 show that copy number can change during sexual recombination because the copy number in these F1 progeny clones does not match copy number in either parent. (D–F) Whole-chromosome alignments between ME49ToxoDB-48 and our Nanopore assemblies at loci harboring tandem gene arrays. Gray boxes with red borders indicate one-to-one mapping regions ≥10,000 bp determined by NUCmer, and orange/blue boxes are as described in Figure 5. Black bars indicate size of select tandem repeats in the ToxoDB and Nanopore assemblies.











