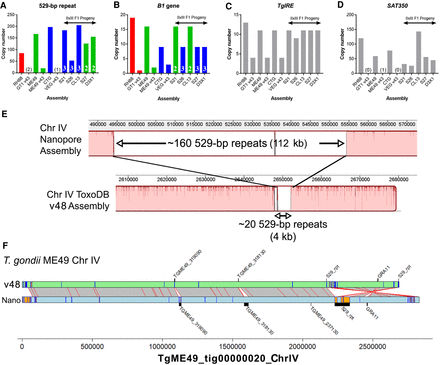
Long-read sequence assemblies precisely resolve canonical repeat sequences and identify additional expansions at gene-harboring loci. (A–D) Estimated copy number for Nanopore assemblies and existing genome assemblies on ToxoDB (“v48”) for T. gondii strain types I, II, and III and II×III F1 progeny. In all cases, Nanopore assemblies identified higher numbers of each repeat locus. In the F1 progeny, B1 gene copy number tracked directly with the genotype (type II or III) at that locus (A), whereas these same F1 progeny harbored unique numbers of 529-bp repeat copies, all of which were not only distinct from their respective genotypes of origin but distinct from one another (B). The TgIRE and SAT350 repeats also were better resolved in our Nanopore assemblies (C,D), although determining genotype of the corresponding region is not possible because these repeats are found at multiple locations throughout the genome. (E) Whole-chromosome alignment focused on the 529-bp repeat region for the v48 ToxoDB assembly (bottom) and our Nanopore-based assembly (top). Expansion of the known genome sequence at this locus in the Nanopore sequence compared with the ToxoDB assembly is clear, as well as consistent with our identification of approximately 140 previously unknown 529-bp repeats in the ME49 genome. (F) Alignment and annotation of repeat sequences of ME49 v48 ToxoDB Chromosome IV and that from our polished Nanopore assembly. Gray bars with red borders indicate mapping regions ≥10,000 bp determined using NUCmer, whereas orange boxes with blue borders indicate tandem repeats with period sizes ≥500 bp and at least two copies. Bars that appear orange are larger than those that are only blue. Incorrect inversion on the right arm of Chromosome IV in the ToxoDB assembly is evident, as is the more accurately resolved 529-bp repeat locus, which was likely a cause for the inversion in standard assemblies from multiple strains.











