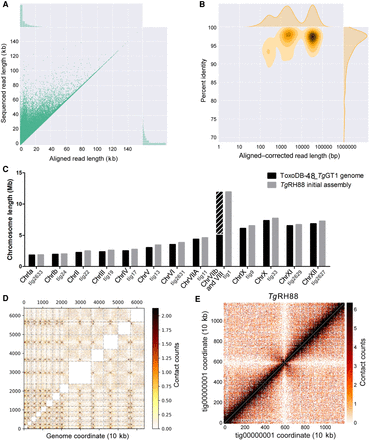
Primary de novo assembly of TgRH88 genome using Nanopore reads revises T. gondii karyotype. (A) Bivariate plot showing a comparison of the aligned read length with the sequenced read length. (B) Bivariate plot showing a comparison of the aligned–corrected read length (log10-transformed) with the percentage identity. In this case, corrected reads refer to the method deployed by Canu using read overlap. (C) Histogram showing comparison of chromosome size between the ToxoDB-48_TgGT1 genome and TgRH88 initial long-read assembly. (D) Inter-chromosomal Hi-C contact-count heat map plotted using the TgRH88 initial long-read assembly sequence showing 13 chromosomes in the assembly. (E) Intra-chromosomal Hi-C contact-count heat map plotted using the sequence of TgRH88_tig00000001 in TgRH88 initial long-read assembly showing no aberrant signal along the contig.











