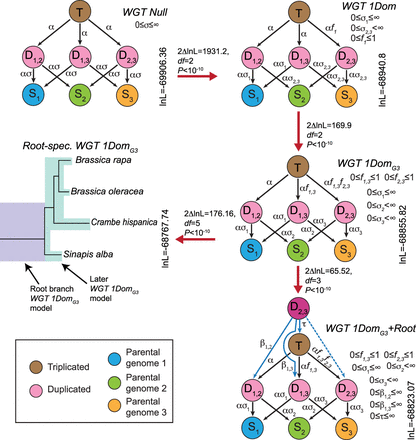
POInT's models for inferring WGT. Five different models of post-WGT evolution and their ln-likelihoods are shown. In each model, the colored circles represent different states. The brown circle represents the triplicated state (T); the pink circles are duplicated states (D1,2, D1,3, and D2,3); the blue, green, and yellow circles are three single-copy states (S1 for the LF subgenome, S2 for the MF1 subgenome, and S3 for the MF2 subgenome). The transition rates between states are shown above the arrows: (α) transition rate from triplicated state to duplicated states; (ασ) transition rates from duplicated states to single-copy states; (f) fractionation parameters; (β and τ) root model parameters. Red arrows connect pairs of models compared using likelihood ratio tests (Methods). In the WGT Null model, transition rates are the same across three subgenomes, modeling the scenario of no biased fractionation. In the WGT 1Dom model with the biased fractionation parameter f1 (0 ≤ f1 ≤ 1), the MF1 and MF2 subgenomes are more fractionated than LF subgenome. In the WGT 1DomG3 model, two fractionation parameters f1,3 and f2,3 were introduced, distinguishing the three subgenomes: MF2 is more fractionated than MF1, and MF1 is more fractionated than LF. The Root-spec. WGT 1DomG3 model is similar to the previous model, but with two sets of parameters, one set for the root branch and the other for the remainder of the branches. The WGT 1DomG3 + Root model is a two-step hexaploidy model created by starting each pillar in an intermediate state D2,3. This state represents the merging of the MF1 and MF2 subgenomes as the first step of the hexaploid formation. The T, D1,2, and D1,3 states represent the second allopolyploidy, with either no prior homoeolog losses (T) or a loss from one of the two MF subgenomes before that event (D1,2, or D1,3).











