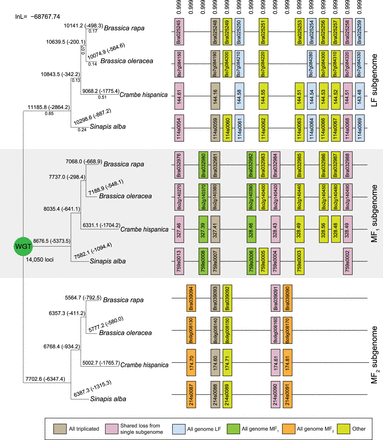
Subgenome assignment and inference of gene loss after the shared WGT in four species. After the WGT, each ancestral locus could potentially expand to three gene copies, but owing to biases in the loss events, the number of surviving genes from the subgenomes are unequal. Our analyses (Results) indicate the presence of a less fractionated (LF) subgenome and two more fractionated ones (MF1 and MF2). These inferences are based on the gene losses observed across four genomes and along the phylogeny depicted. Shown here is a window of 16 post-WGT loci (of the total 14,050 such loci) in four species that share the WGT: Brassica rapa, Brassica oleracea, Crambe hispanica, and Sinapis alba. Each pillar corresponds to an ancestral locus, and the boxes represent extant genes. Pairs of genes are connected by lines if they are genomic neighbors (e.g., in synteny). The numbers above each pillar are the posterior probabilities assigned to this combination of orthology relationships relative to the other (3!)4−1 = 1295 possible orthology states. The numbers above each branch of the tree give the number of genes in each subgenome surviving to that point, with the number of gene losses in parentheses. The gene loss inferences made by POInT are probabilistic: because some gene losses cannot be definitively assigned to a single branch, the resulting loss estimates are not integers. The numbers below the branches in the first subtree are POInT's branch length estimates (αt).











