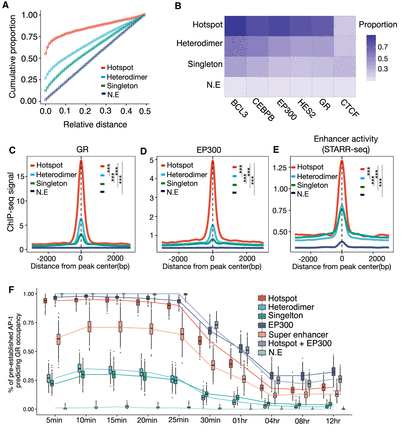
Genomic landscape of various AP-1 binding modes in glucocorticoid response. (A) Cumulative distribution of relative distance of AP-1 binding classes to the nearest genomic loci occupied by GR. (B) Heat map showing the means of the Jaccard index of overlaps between AP-1 classes and genomic occupancies of TFs. The means are based on the distribution of the indexes from bootstrapped replicates of genomic regions that matched in number. (C–E) Spatial distribution of dex-responsive TFs and histone marks for distinct AP-1 binding classes: (C) glucocorticoid receptor (GR); (D) coactivator EP300; (E) enhancer activity defined by genome self-transcribing active regulatory region-seq (STARR-seq). P-values were calculated using a two-sided Student's t-test based on ChIP-seq signal intensity within 1 kb centered on each peak. (***) P < 0.001. (F) The distribution of proportions of dynamic GR bindings explained by pre-established factors including AP-1s, EP300, and super-enhancers at a pre-dex time point.











