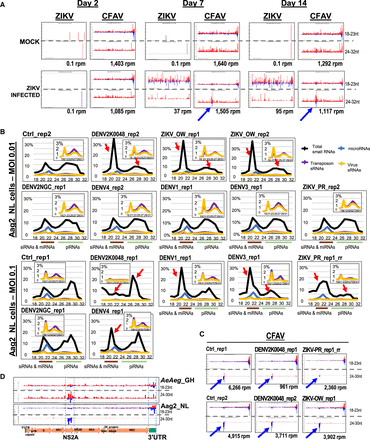
Small RNA cross talk in Aedes aegypti (AeAeg) during flavivirus infections. (A) Reanalysis of ZIKV and CFAV small RNAs from AeAeg females as sequenced from Saldaña et al. (2017). The blue arrow notes emergence of new piRNAs from CFAV after active replication of ZIKV small RNAs. The x-axis gives the coordinates of the virus sequence; the y-axis is the autoscaled read frequency. The total small RNA normalized counts are below each plot. (B) Small RNA length distributions as a proportion of the small RNA library. The inset graph zooms in on the modest proportions of viral and transposons small RNAs. Red arrows point to the significant change from the normal proportion of small RNAs in control cells. (C) Counts and small RNA profiles from CFAV in control and infected Aag2-NL cells. Blue arrows point to preexisting group of negative strand piRNAs potentially because of multiple preexisting viruses replicating and generating small RNAs in Aag2-NL cells. (D) The regions generating notable piRNAs and siRNAs from CFAV in mosquitoes and Aag2 cells are the NS2A gene and 3′ UTR.











