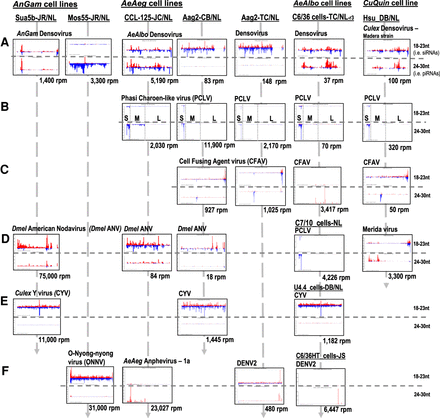
Multiple arboviruses persistently infect mosquito cell cultures and generate arboviral small RNAs. Profiles of viral small RNAs in cell culture lines from AnGam, CuQuin, AeAeg and AeAlbo. Reads per million (rpm) numbers are totals of the siRNA-length and piRNA-length small RNAs that come from the plus strand in red and minus strand in blue. The x-axis gives the coordinates of the virus sequence, and the y-axis is the autoscaled read frequency. The total small RNA normalized counts are below each plot. The suffix to sample names is the initials of the laboratory investigator where the sample was originally obtained: (JR/NL) Jason Rasgon to Nelson Lau; (DB) Doug Brackney; (JC) John Connor; (CB) Carol Blair; (TC) Tonya Colpitts; (JS) Juan Salas-Benito. The S, M, and L segments of the Phasi Charoen-Like virus (PCLV) are marked on these coverage plots. (A) Various species densoviruses. (B) Phasi Charoen-like virus. (C) Cell fusing agent virus. (D) Drosophila American nodavirus and two other cells with PCLV and Merida virus. (E) Culex Y virus. (F) Alphaviruses and flaviviruses.











