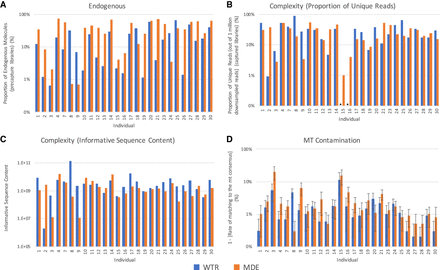
Sample quality. A comparison of the quality of data produced by WTR (Whole Tooth Root) and MDE (Minimally Destructive Extraction) methods in samples that passed quality filtering. (A) The proportion of endogenous molecules in data obtained via shotgun sequencing. (B) The complexity of each sample, as measured by the proportion of unique reads out of 1,000,000 reads sequenced. Asterisks indicate that the total number of unique reads sequenced was below 1,000,000 for the specified sample, and therefore complexity estimates could not be generated. (C) The complexity of each sample, as measured by informative sequence content. (D) The rate of contamination is compared by considering the rate of matching to mitochondrial consensus sequence. Error bars indicate the 95% confidence interval. Only samples that passed quality screening are shown. Plots showing comparisons with samples generated using Method P are shown in Supplemental Figure S2.











