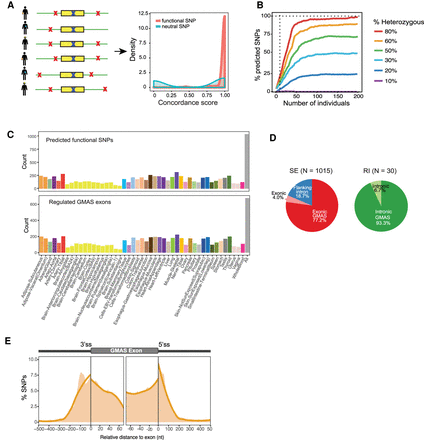
Prediction of functional SNPs for GMAS events. (A) Functional SNPs are predicted by considering candidate SNPs (red crosses) in the vicinity of a GMAS exon, including the tag SNP itself (blue crosses). Concordance among the allelic ratios of the tag SNP in all samples is calculated as described in Methods (with hypothetical distributions shown). (B) The percentage of SNPs predicted given the number of individuals in the simulated testing cohort (Methods). Different percentages of individuals with the heterozygous genotype were simulated. Vertical dotted line marks 10 individuals. (C) Top: Number of predicted functional SNPs per tissue. Bottom: Number of GMAS exons with predicted functional SNPs per tissue. The rightmost bar (All) corresponds to predictions made by pooling samples from all tissues. (D) Left pie chart: predicted functional SNPs in the exonic or intronic regions of SE (skipped exons). Exonic GMAS: the functional SNP is also the exonic GMAS tag SNP. The rest of the functional SNPs were classified into the “Exonic” or “Flanking intron” group. Right pie chart: for retained introns (RI). Intronic GMAS: the functional SNP is also the GMAS tag SNP. N refers to the number of functional SNPs for each group. No functional SNPs were predicted for alternative 5′ or 3′ ss exons. (E) Densities of predicted functional SNPs near the exon-intron boundaries of their associated GMAS exons (SEs only). The number of functional SNPs was normalized by the total number of testable SNPs at each nucleotide position. Orange curve is the fitted trend line of the shaded area that represents the SNP density at single-nucleotide resolution.











