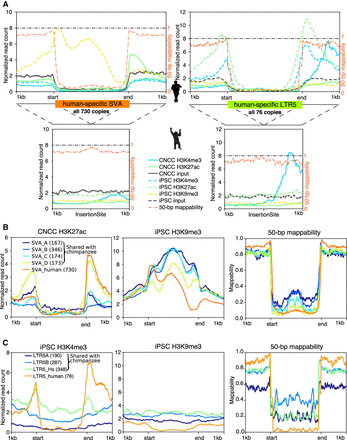
ChIP-seq read count, normalized to reads per genomic content, surrounding repetitive TE insertions reveals putative hidden CREs. (A) ChIP-seq signal profiles and 50-bp mappability over all human-specific SVA and LTR5 insertions in the human genome (top) and their orthologous preinsertion sites in the chimpanzee genome (bottom), with 1-kb flanking regions. (B) Profile of CNCC H3K27ac ChIP-seq, iPSC H3K9me3 ChIP-seq, and 50-bp mappability around SVA insertions in the human genome, with 1-kb flanking regions. SVA subfamilies are distinguished by color with copy numbers inside parenthesis. (C) Profile of iPSC H3K4me3 ChIP-seq, iPSC H3K9me3 ChIP-seq, and 50-bp mappability around LTR5 insertions in the human genome, with 1-kb flanking regions. LTR5 subfamilies are distinguished by color with copy numbers labeled.











