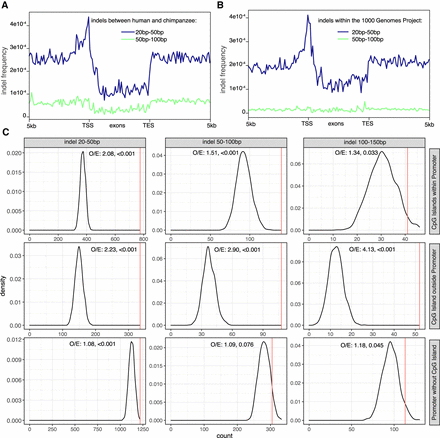
Enrichment of 20- to 50-bp indels in promoters/CGI. (A) Frequency of human–chimpanzee indels of sizes 20–50 bp and 50–100 bp in and around annotated human genes. Metaplots display genes with introns removed and 5-kb flanking regions surrounding the transcription start site and end site. (B) Frequency of human population indels found in The 1000 Genomes Project around the same promoter regions described in A. (C) Comparing the observed number of indels intersecting with CpG island within promoters, CpG island outside of promoters and promoters without CpG island with the same intersections between randomly shuffled indels and the three CpG island or promoter regions. All indels were shuffled 1000 times. The density distributions of the numbers of shuffled indel intersections are illustrated by black lines. The observed numbers are indicated with a red vertical line with the observation/expectation ratio (O/E) and P-value at the top of each graph.











