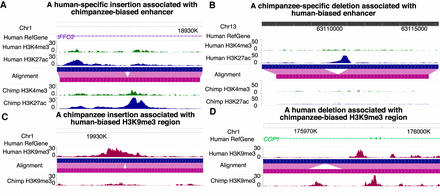
Examples of indels associated with different CREs on the WashU Epigenome Browser. Human–chimpanzee track represents pairwise alignment between the human (blue) and chimpanzee (pink) genomes. (A) A human-specific insertion associated with a chimpanzee-biased enhancer. (B) A chimpanzee-specific deletion associated with a human-biased enhancer. (C) A chimpanzee-specific insertion associated with a human-biased H3K9me3 heterochromatin region. (D) A human-specific deletion associated with a chimpanzee-biased H3K9me3 heterochromatin region.











