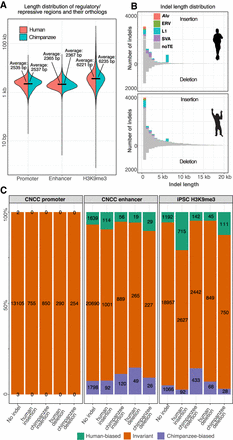
All putative regulatory and repressed regions (RRRs) and indels between human and chimpanzee and the number of overlaps between them. (A) Length distribution of all putative RRRs and their orthologs. Violin plots show the length of putative promoter, enhancer, and H3K9me3-repressed regions. For regions called in only one species, the length of their syntenic regions are plotted in the other species. Size in the human genome is shown on the left; size in the chimpanzee on the right. The average length of each distribution is marked and labeled. (B) Size distribution of indels between human and chimpanzee. The number of indels in each lineage is plotted in a back-to-back histogram with indel length on the x-axis and the number of indels of different lengths on the y-axis. Colors distinguish indels based on TE classification. (noTE) Not derived from a TE insertion. (C) The number of lineage-biased/invariant putative CNCC promoters, CNCC enhancers, and iPSC H3K9me3 heterochromatin regions with or without indel association. Regions were separated into those without an indel or with one of the four types of indels. Colors distinguish putative RRR invariant between the two species or biased in either lineage. The percentage of each category is displayed in a stacked histogram with the number of occurrences labeled.











