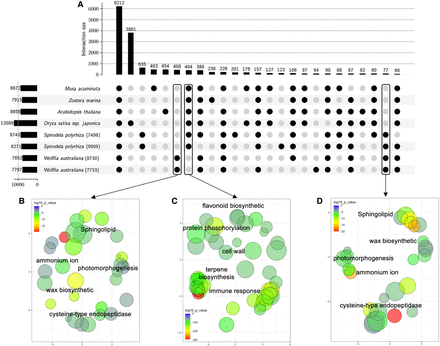
Wolffia orthogroups (OGs) analysis reveals significant Gene Ontology (GO) terms relating to growth and defense. (A) An upset plot showing the orthogroups (OGs) present (black circles) or missing (gray circle) across the four duckweed genomes (sp9509, sp7498, wa7733, and wa8730), model species (Arabidopsis thaliana, Oryza sativa ssp. Japonica), and two non-grass monocots (Musa acuminata, Zostera marina). Boxes indicate Wolffia-specific, Wolffia-missing, and duckweed-specific genes. (B) Significant (FDR < 0.05) GO terms derived from Wolffia-specific OGs (A) were plotted in two-dimensional semantic space by multidimensional scaling (MDS) with the color reflecting significance (P-value) and circle size reflecting GO frequency. (C) GO terms from Wolffia missing OGs. (D) GO terms from duckweed-specific OGs. All GO terms were summarized using semantic similarity (SimRel), and P-values from overrepresentation with REVIGO.











