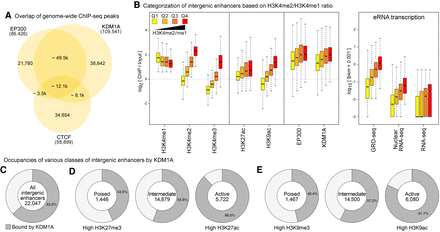
KDM1A occupies a large fraction of enhancers in mESCs. (A) Overlap of binding sites of EP300, CTCF, and KDM1A in mESCs. (B) Intergenic enhancers were divided into quartiles (Q1–Q4) based on the enrichment of H3K4me2 relative to H3K4me1 (left panel). Box plots show the enrichment of the indicated histone modifications, KDM1A, and EP300, as measured using ChIP-seq and eRNA levels (GRO-seq, nuclear RNA-seq, and RNA-seq) at each quartile of the intergenic enhancers. Levels of KDM1A show positive correlations with increases in H3K4me2 and eRNA expression from Q1 to Q4. In all figures, the bottom and top boxes signify the second and third quartiles, respectively, and the middle band represents the median of the population. Whiskers represent 1.5 times the interquartile range (IQR), and the notch represents the 95% confidence interval of the median. (C) The percentage of intergenic enhancers with KDM1A peaks. (D,E) Active, poised, and intermediate enhancers were classified based on the enrichment of either trimethylation or acetylation of H3K27 (D) or H3K9 (E). KDM1A occupancy at enhancers increases with higher enhancer activity.











