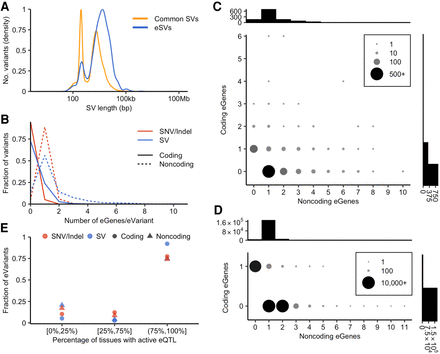
Features of SV-eQTLs. (A) Size distribution of eSVs compared with all common SVs. (B) Distribution of the number of eGenes per eVariant for SVs compared with SNVs and indels. “Coding” eGenes refer to eGenes whose exons are intersected by the associated eVariant, and “noncoding” eGenes are not intersected by the associated eVariant. Counts are shown for every eVariant; thus, eVariants with zero coding or zero noncoding eGenes are included in the distributions. (C,D) The number of eVariants, as shown by dot size and color, with the indicated combination of coding and noncoding eGenes, as defined above. Shown for SVs (C) and SNV/indels (D), with histograms showing the total number of eVariants, with the indicated number of associated coding or noncoding eGenes above the y- and x-axes, respectively. (E) Distribution of tissue specificity of eQTLs across tissues as evaluated by METASOFT, separated into the lowest quartile, middle two quartiles, and top quartile, for eQTLs in which the activity status is known in at least 43 of 48 evaluated tissues. The points indicate the fraction of SV-eQTLs or SNV- and indel-eQTLs that are active (m > 0.9) in the proportion of tissues indicated on the x-axis.











