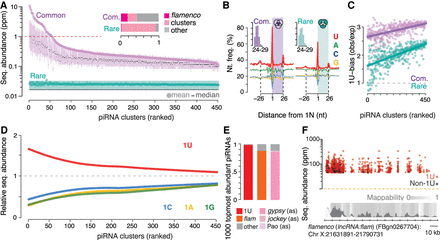
Precursor and processing preferences determine piRNA sequence abundance. (A) A small number of top-ranked piRNA clusters (Supplemental Table S3), led by flamenco (lncRNA:flam), produces common sequences (purple) with an average abundance of more than one read per million (red dotted line). Clusters were ranked by the mean abundance of common sequences (Supplemental Table S3). The sequence abundance of common and rare Piwi-piRNAs for each cluster is shown (mean, median, 25th–75th percentile; n = 3). Inset: The fraction of common and rare Piwi-piRNAs that originate from flamenco (lncRNA:flam) and other piRNA clusters. (B) Common and rare Piwi-piRNAs are generated by the Zuc-processor. Metagene analyses reveal the phased preference of the Zuc-processor for uridine (U) in the first position of the observed piRNAs (colored box) as well as in the first position of proceeding and preceding piRNAs. Inset: Length distribution of common and rare piRNAs in nucleotides (nt). (C) Common piRNAs have a stronger 1U-bias than rare piRNAs from the same cluster. The bias for uridine (1U-bias) in the first position—observed over expected 1U frequencies (obs/exp)—of Piwi-piRNAs (clusters ranked as in A and D). (D) Uridine in the first position (1U) is indicative of higher sequence abundance. The mean abundance of piRNAs that start with a uridine (1U) was compared to that of piRNAs that start with either adenosine (1A), guanosine (1G) or cytidine (1C). The mean abundance of 1U-, 1A-, 1G-, and 1C-sequences relative to the mean abundance of all sequences from the same cluster (relative sequence abundance) is shown (clusters ranked as in A and C). (E,F), Most of the 1000 topmost abundant Piwi-piRNAs exhibit a 1U, originate from flamenco (lncRNA:flam) (flam) and show antisense-complementarity (as) to gypsy endogenous retroviruses. Annotated fractions (E) and genomic position within flamenco (lncRNA:flam) (F). Multimapping sequences are represented with all possible coordinates within flamenco (lncRNA:flam) (F).











