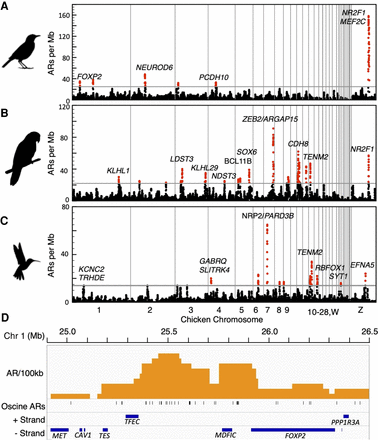
Accelerated region distributions. (A–C) AR hotspot plots in genomes for the three vocal learning lineages using chicken as the reference for chromosome location and annotations. Each dot indicates the number of ARs in a 1-Mb window. Windows overlap and are offset from one another by 100 kb. Regions with AR density higher than random are “hotspots” and labeled in red dots. Gray vertical lines demark chromosome boundaries; chromosomes are arranged 1–28, W, Z. Genes in acceleration hotspots with known speech or neurodevelopmental functions and/or associated with convergent hotspots in vocal learners are indicated in each plot. (D) An enlarged view of the distribution of ARs per 100 kb in the FOXP2 containing acceleration hotspot. Genes in the hotspot are arrayed by strand (+ or −). FOXP2 is the only gene in this region with predicted vocalization behavior function based on coexpression (Z = 3.48). Silhouettes by Anthony Caravaggi and Ferran Sayol.











