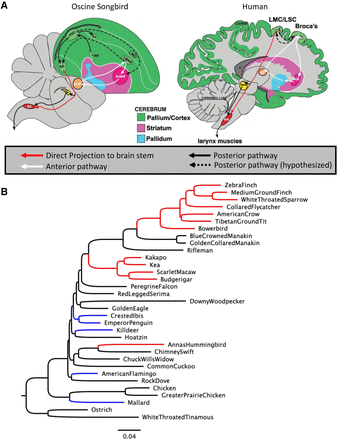
Convergent lineages. (A) Diagram of the vocal learning–related brain regions and circuits in oscine songbirds and human. Both species are characterized by anterior learning (white arrows) and posterior production (solid black arrows) pathways, including a direct neuronal projection (red arrow) from the forebrain to the brain stem. (B) A phylogeny of the taxa in this study using the topology of Jarvis et al. (2014) with branch lengths inferred from fourfold degenerate sites. Branches are colored by convergent phenotype: Red: vocal learners; blue: waterbirds, which are used as a biological control for some analyses; black: species that do not belong to either of these groups. Abbreviations: Songbird brain: Area X, Area X of the striatum; AV, nucleus avalanche; DLM, dorsolateral nucleus of the medial thalamus; DM, dorsal medial nucleus of the midbrain; HVC, high vocal center; LMAN, lateral magnocellular nucleus of the anterior nidopallium; LMO, lateral oval nucleus of the mesopallium; Nif, interfacial nucleus of the nidopallium; RA, robust arcopallium; XII, bird twelfth nerve nucleus. Human brain: Am, nucleus ambiguous; ASt, anterior striatum; aT, anterior thalamus; LMC, laryngeal motor cortex; LSC, laryngeal somatosenosry cortex; PAG, periaqueductal gray.











