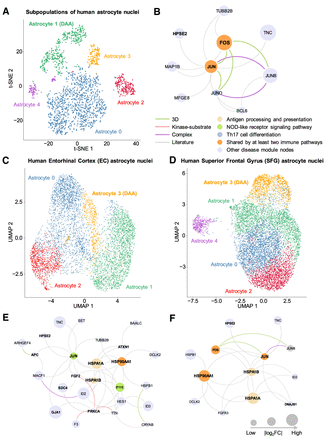
Discovery of DAA-specific molecular networks from single-nucleus RNA sequencing data of human brains with AD. (A) T-distributed stochastic neighbor embedding (t-SNE) plot of clustering 2119 astrocyte nuclei between AD patients and healthy controls. (B) An identified DAA-specific molecular network contains 16 protein–protein interactions (PPIs) connecting 10 gene products (proteins). (C) UMAP plot for 5599 astrocyte nuclei clustering analysis of brain entorhinal cortex (EC) regions among AD patients with different Braak stages. (D) UMAP plot of clustering 8348 astrocyte nuclei for brain superior frontal gyrus (SFG) regions among AD patients with different Braak stages. (E) An identified DAA-specific molecular network containing 43 protein–protein interactions (PPIs) connecting 26 gene products (proteins) for EC. (F) An identified DAA-specific molecular network containing 22 PPIs connecting 13 genes/proteins for SFG. Node sizes are proportional to their corresponding |log2FC|. Node color is coded by known immune pathways from the KEGG database. Edge color is coded by experimental evidences of PPIs. Key immune modulators related to AD are highlighted by bold text.











