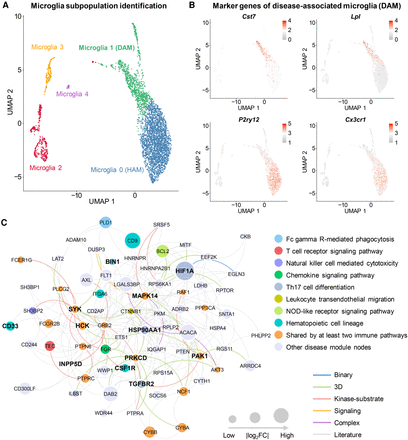
Discovery of DAM-specific molecular networks for the transgenic mouse model of AD. (A) Uniform manifold approximation and projection (UMAP) plot of clustering 4389 microglia cells: the blue cluster denotes the homeostasis-associated microglia (HAM), and the green cluster denotes the DAM. (B) Expression levels (heatmap) of representative marker genes (up-regulation in DAM: Cst7 and Lpl; and down-regulation in DAM: P2ry12 and Cx3cr1) in different microglia subclusters. (C) A predicted DAM-specific molecular network contains 227 protein–protein interactions (PPIs) connecting 72 proteins. Node sizes are proportional to their corresponding |log2FC| during differential expression analysis. (FC) Fold-change. Node (gene/protein) color is coded by known immune pathways from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Edge color is coded by known experimental evidences of PPIs (Methods). Key immune modulators related to AD are highlighted by bold text.











