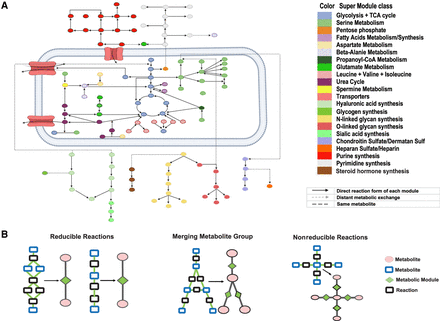
Reduced and reconstructed human metabolic map. (A) Collected human metabolic modules and supermodule classes. (B) Examples of how the network motifs in the metabolic map are simplified into metabolic modules, where the reactions and metabolites are represented by black and blue rectangles, and modules and metabolites are colored by green and pink. Chainlike reactions can be directly simplified; a complicated module connected by multiple branches can be shrunk into one point linked with the multiple in/out branches; and complicated intersections cannot be simplified.











