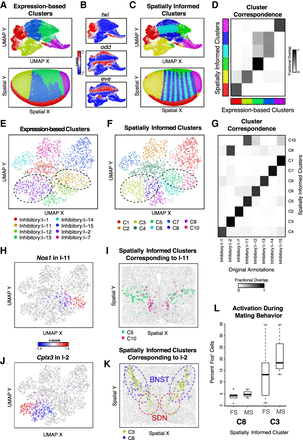
Spatially informed clustering distinguishes spatially distinct subpopulations of cells. (A) Expression-based clustering of 3035 stage 6 D. melanogaster embryo cells with 84 marker genes by aligned ISH identifies approximately five transcriptionally distinct clusters. (Top) UMAP embedding colored by identified cluster annotations. (Bottom) Spatial coordinates colored by identified cluster annotations. (B) Expression of select marker genes on the UMAP embedding with red denoting high expression and blue denoting low expression. (C) Spatially informed clustering splits expression-based clusters in a spatially coherent manner. (Top) Again, UMAP embedding colored by identified spatially informed cluster annotations. (Bottom) Spatial coordinates colored by identified spatially informed cluster annotations. (D) Correspondence between expression-based clusters in A and spatially informed clusters in C highlights high correspondence between most clusters with the exception of one cluster being split into two. (E) UMAP embedding of populous inhibitory neuronal subtypes in one posterior preoptic tissue section from one animal measured using MERFISH, where each point is a cell colored by the original subtype annotations. (F) Same UMAP embedding as E in which each point is a cell colored by the spatially informed clustering annotation. Black dashed lines highlight clusters that have now split. (G) Correspondence between expression-based clusters in E and spatially informed clusters in F highlights high correspondence between most clusters with the exception of cells originally annotated as I-2 and I-11 now being split into two. (H) Same UMAP embedding as E in which each point is a cell colored by Nos1expression for cells originally annotated as I-11. (I) Spatial location of cells within the tissue colored by their spatially informed cluster assignment for cells originally annotated as I-11. (J) Same UMAP embedding as E, in which each point is a cell colored by Cplx3 expression for cells originally annotated as I-2. (K) Spatial location of cells within the tissue colored by their spatially informed cluster assignment for cells originally annotated as I-2. Regions corresponding to the BNST and SDN are highlighted with blue and red dashed lines, respectively. Representative slice in representative animal shown. (L) Percentage of activated cells based on Fos expression during female (FS) and male (MS) sexual behavior for spatially informed clusters C3 and C8 originally annotated as I-2. Boxes in the box plot denote the median values and inner quartile ranges (IQR), and whiskers denote 1.5 × IQR with additional outliers represented as points.











