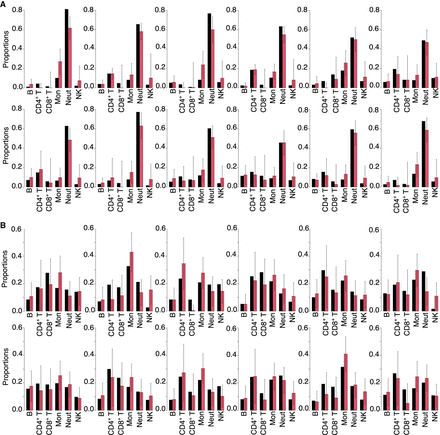
RNA-Sieve results with confidence intervals for whole blood bulk samples with known cell type proportions. Inferred cell type proportions in deconvolutions using PBMC references and whole blood bulks as described in “Validation with real bulk RNA-seq data”: (A) Newman et al. (2019) data; (B) Monaco et al. (2019) data. True proportions as estimated by flow cytometry are in black, and RNA-Sieve's inferred proportions are in red. The black error bars on inferred proportions show the marginal 95% confidence intervals as produced by RNA-Sieve. Confidence intervals capture true proportions 90.3% and 95.8% of the time in the respective scenarios. See “Confidence intervals” for mathematical details.











