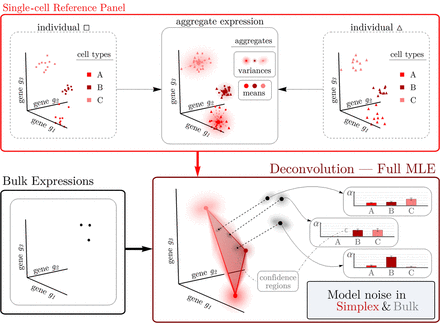
The RNA-Sieve pipeline. After applying a filtering procedure to scRNA-seq data, RNA-Sieve builds reference matrices for the mean and variance of expression for each gene across cell types. Using these estimates and bulk RNA-seq data, it performs joint deconvolution via maximum likelihood estimation by expressly modeling noise both in the reference and bulk data, yielding cell type proportion estimates and confidence regions for each sample.











