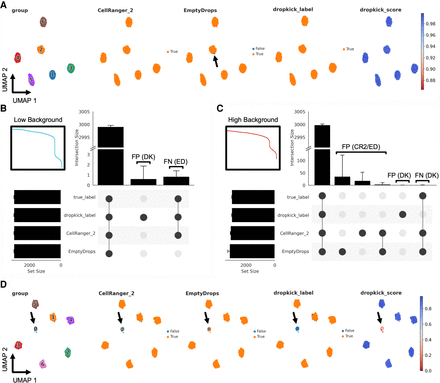
Evaluating dropkick filtering performance with synthetic data. (A) UMAP embedding of all barcodes kept by dropkick_label, CellRanger_2, and EmptyDrops for an example low-background simulation. Points colored by each of the three filtering labels, as well as ground-truth clusters determined by the simulation and dropkick score (cell probability). Arrow highlights a single false-negative (FN) barcode in the EmptyDrops label set for this replicate. (B) UpSet plot showing mean size of shared barcode sets across dropkick_label, CellRanger_2, EmptyDrops, and true labels for 10 simulations. Error bars, SD. Unique sets show false-positive (FP) barcodes labeled by dropkick and FN barcodes excluded by EmptyDrops. Inset shows log-rank representation of the low-background simulation in A. (C) Same as in B, for 10 high-background simulations. Inset shows log-rank representation of the high-background simulation in D. (D) Same as in A, for an example high-background simulation. Arrow highlights cluster 0, designated as “empty droplets” by simulation (see Methods: Synthetic scRNA-seq data simulation).











