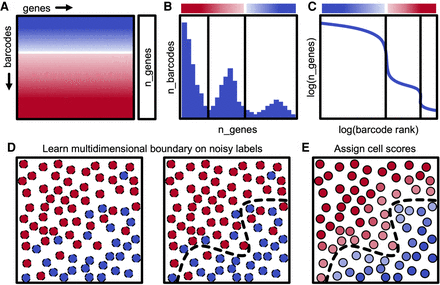
Description of dropkick filtering method. (A) Diagram of scRNA-seq counts matrix with initial cell confidence for each barcode based solely on total genes detected (n_genes), depicted by color (red, empty droplet; blue, real cell). (B) Histogram showing the distribution of barcodes by their n_genes value. Black lines indicate automated thresholds for training the dropkick model. (C) log(n_genes) versus log(rank) representation of barcode distribution as in dropkick QC report (Fig. 1A). Thresholds from B are superimposed. (D) Thresholds in heuristic space (B,C) are used to define initial training labels for logistic regression. (E) dropkick chooses an optimal regularization strength through cross-validation and then assigns cell probabilities and labels to all barcodes using the trained model in gene space.











