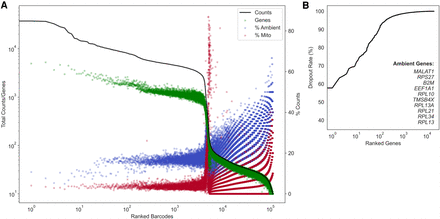
Evaluating data set quality with the dropkick QC module. (A) Profile of total counts (black trace) and genes (green points) detected per ranked barcode in the 4000 pan–T cell data set (10x Genomics). Percentage of mitochondrial (red) and ambient (blue) reads for each barcode included to denote quality along data set profile. (B) Profile of dropout rate per ranked gene. Ambient genes are identified by dropkick and used to calculate ambient percentage in A.











