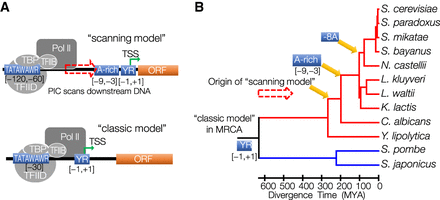
The origin and evolution of the scanning model of transcription initiation mechanism. (A) Schematic illustration of two distinct mechanisms of transcription initiation: classic model and scanning model. Both initiation mechanisms share a strong preference for PyPu dinucleotides at the [−1,+1] sites. In the classic model, transcription initiates at ∼30 bp downstream from the TATA box, where PIC is assembled. In contrast, the PIC in scanning-model species scans DNA downstream from the TATA box for TSSs with favorable genomic context, which includes a PyPu and an A-rich region immediately upstream of it. (B) Schematic illustration of the origin of the scanning model and associated genetic innovations during the evolution of budding yeasts. The most recent common ancestor (MRCA) of the budding yeasts and fission yeasts was the classic-model species. The scanning model originated after the split between Y. lipolytica from the other budding yeasts. An A-rich region in the region from −9 to −3 bp upstream of the TSS appeared during the evolution of scanning-model species after the divergence of C. albicans. The gaining of specific −8A preference occurred in ancestral WGD species.











