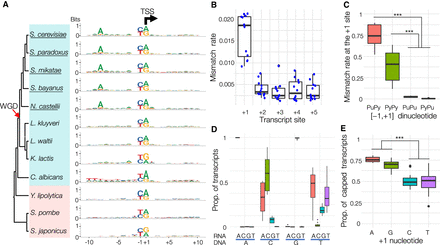
Functional roles of PyPu dinucleotide in transcription initiation. (A) Consensus sequences of core promoters in the 12 yeast species. The sequence logo was generated using sequences from −10 to +10 bp surrounding the dominant TSS of all core promoters in each species (Supplemental Dataset S13). The black arrow indicates the TSS position (the +1 site) and the transcription direction. The red arrow indicates the occurrence of WGD, and the names of WGD species are underlined. (B) Boxplot of the distributions of mismatched rates at the first five sites of transcripts in the 12 species. Each blue dot represents the mismatch rate at each site in a species. (C) Mismatch rates in transcripts initiated from different −1/+1 dinucleotides: PuPy, PyPy, PuPu, and PyPu. (*) P < 0.01; (**) P < 0.001; (***) P = 0. (D) Proportion of each type of nucleotides added by Pol II at the +1 site of RNA transcripts. On the x-axis, the type of recruited nucleotides (RNA) is shown above the blue lines, and the nucleotides on the sense strand (DNA) are shown under the blue lines. (E) Boxplot illustrates proportions of transcripts with a detected G-cap at the 5′ end among transcripts with different starting nucleotides in the 12 yeast species.











