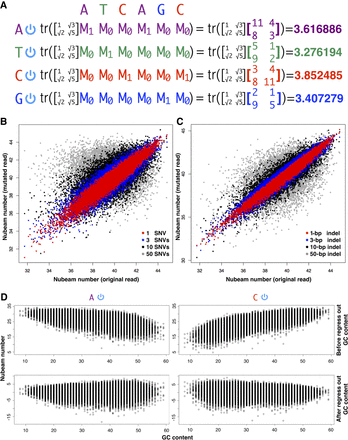
Nubeam assigns numbers to reads. (A) Illustration of how to obtain Nubeam quadruplet for a read. Convert a read to four binary sequences (indicated by the on/off power symbol), turn each binary sequence into a product matrix, and obtain a number from each product matrix. (B,C) Similar binary sequences produce similar numbers. For each simulated binary sequence of length 100, we obtained sequences with 1 or 3 or 10 or 50 random SNVs, or sequences with 1- or 3- or 10- or 50-bp indels, and compared the Nubeam numbers of original sequences with those of mutant sequences. (D) Regressing out GC content from Nubeam numbers of binary sequences; data comes from mapped reads of HMP sample SRS019215. Left: With A as reference (T as reference is similar). Right: With C as reference (G as reference is similar).











