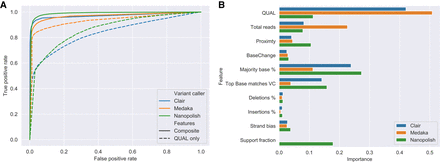
Random forest–based variant filtering using Nanopolish, Medaka, and Clair. (A) Receiver operating characteristic (ROC) curve for random forest classifier using different features including Quality (QUAL only, dashed line) and a composite selection of input features (Composite, solid line) for Nanopolish (green), Medaka (orange), and Clair (blue). AUC for each variant caller: Nanopolish 0.86 to 0.98, Medaka 0.93 to 0.97, Clair 0.84 to 0.97, using QUAL and Composite features, respectively. (B) Bar chart of feature importance for composite selection of features used to train the classifier.











