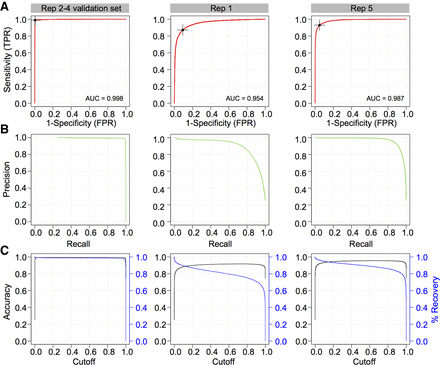
Performance of 2D convolutional neural network barcode classifier. (A) Receiving operator characteristic (ROC) analysis and area under the curve (AUC) metrics of the final model on three evaluation sets: (1) Replicates 2–4 validation set (left column), which was generated from the same sequencing runs used to train the model but were withheld from training; (2) Replicate 1 set (middle column), composed of reads generated using the RNA001 library kit; and (3) Replicate 5 set (right column), derived from an independent sequencing run using the RNA002 kit. Optimal Youlden index (J statistic) is marked as a black cross on the ROC curve. (B) The associated precision recall curves on the three test sets. (C) Accuracy (black) and percentage of reads recovered (blue) in function of the scoring threshold (cut-off) emitted by the trained model, for three different data sets presented in A.











