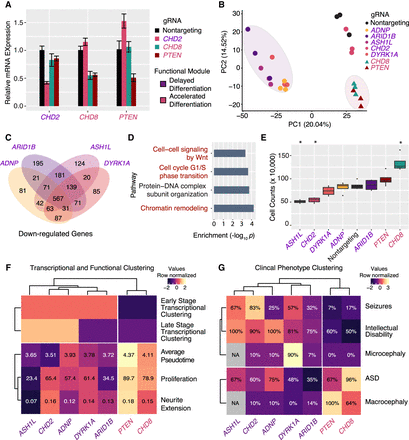
CRISPR repression in iPSC neural progenitor cells confirms modules of ASD genes and transcriptional convergence at cell-cycle dysregulation. Experimental clustering of functional ASD gene modules matches clustering by clinical phenotypes. (A) Efficient dCas9-KRAB repression of individual target genes using the designated gRNAs in iPSC-NPCs. n = 3 biological replicates for all qPCR experiments. Values, mean ± SEM. (B) Clustering of RNA-seq profiles by principal component analysis reveals clustering of “delayed differentiation” and “accelerated differentiation” module ASD genes. (C) “Delayed differentiation” module gene repression elicits strongly overlapping transcriptional consequences. (D) Gene Ontology analysis of down-regulated genes shows enrichment for chromatin remodeling and cell-cycle genes. (E) Cellular proliferation measured by cell number after individual gene repression reveals significant decreases or increases. (*) P < 0.01, n = 4). (F) Integrating transcriptional and functional assays reveals and refines two functionally convergent modules of ASD genes. (G) Clinical phenotype data reveal the same two modules of ASD genes. (%) Prevalence of phenotype (percentage) in individuals from Stessman et al. (2017).











