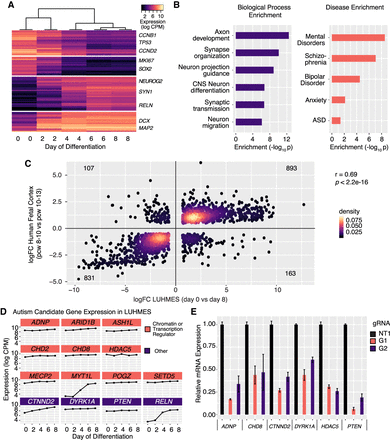
LUHMES are a tractable, disease-relevant model of human neuronal differentiation amenable to perturbation. (A) Hierarchical clustering of bulk RNA-seq time course expression data indicates rapid and reproducible neuronal differentiation. Two replicates for each time point were performed. (B) Genes induced during LUHMES differentiation are enriched for relevant biological processes (left) and neurological disorders (right). (C) Differentially expressed genes during LUHMES differentiation are highly correlated with transcriptional changes that occur during early human fetal corticogenesis (Pearson's rho = 0.69, P = 2.2 × 1016). (logFC) log2 fold-change of differential expression between indicated time points; (pcw) postconception week. (D) High-confidence autism-causing genes, selected for perturbation experiments, are highly expressed at baseline or are increasingly expressed in LUHMES during differentiation and were selected for roles in transcriptional regulation. (E) Efficient dCas9-KRAB repression of individual target genes using the designated guide RNAs. n = 3 biological replicates for all qPCR experiments. Values represent mean ± SEM. (NT1) Nontargeting control gRNA; (G1) gRNA 1; (G2) gRNA 2.











