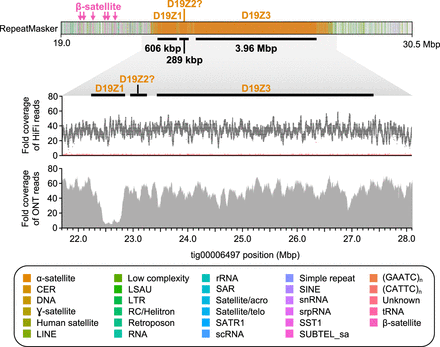
HiCanu assembly of the CHM13 Chromosome 19 centromere. RepeatMasker (Smit et al. 2013) of tig00006497 reveals three α-satellite HOR arrays that reside within the Chromosome 19 centromere (D19Z1, D19Z2?, and D19Z3; marked with black bars). These HOR arrays are 606 kbp, 289 kbp, and 3.96 Mbp in length, respectively, and are composed of a 13-mer, a complex higher-order HOR, and a dimeric HOR unit, respectively. The HOR repeat underlying D19Z2 shares limited sequence identity with the pG-A16 repeat previously described (Hulsebos et al. 1988; Choo et al. 1991; Finelli et al. 1996) and, therefore, is designated with a question mark. The α-satellite HOR arrays have relatively uniform coverage of HiFi and ultralong Oxford Nanopore data, except for a drop in Oxford Nanopore sequencing coverage over the D19Z1 array, which may be owing to a misassembly, read mismapping, or biases in sequencing. The HiFi coverage plot shows fold coverage of the most common base (black) and the second most common base (red).











