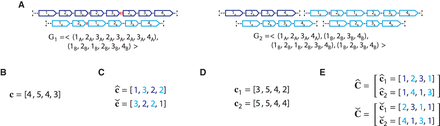
Ambiguity and errors in inferring segment copy number (SCN) profiles for a heterogeneous sample S = (G1, G2) under different assumptions about the sample composition. (A) A two-genome proper sample S = (G1, G2): each genome Gi ∈ S is depicted as collections of adjacent blocks (top), and the corresponding sequences of signed blocks (bottom). (B) The copy number profile c = [c1, c2, c3, c4] inferred under the assumption that the sample is homogeneous (i.e., comprised of a single derived genome) and the reference
genome is haploid (i.e., each segment has only a single haplotype in the reference). Each value cj is the weighted average of the sums of haplotype-specific (or allele-specific) copy numbers 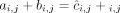 over the genomes Gi ∈ S. (C) Allele-specific copy number profiles
over the genomes Gi ∈ S. (C) Allele-specific copy number profiles 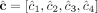 and
and 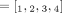 inferred under the assumption that the sample is homogeneous and the reference genome is diploid (i.e., each segment has two haplotypes labeled A and B). Here, the entries
inferred under the assumption that the sample is homogeneous and the reference genome is diploid (i.e., each segment has two haplotypes labeled A and B). Here, the entries 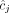 and
and  for segment j are averages
for segment j are averages 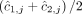 and
and 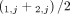 of genome- and allele-specific copy number values. Note that the vectors
of genome- and allele-specific copy number values. Note that the vectors  and č do not preserve the true A/B label of each allele: dark blue are true counts of allele A and light blue are true counts
of allele B. Here, segments 2 and 4 are flipped. (D) Genome-specific copy number profiles c1 = [c1,1, c1,2, c1,3, c1,4] and c2 = [c2,1, c2,2, c2,3, c2,4] inferred under the assumption that the sample is heterogeneous, but the reference genome is haploid. Here, the entry ci,j for a segment j and genome Gi is the sum
and č do not preserve the true A/B label of each allele: dark blue are true counts of allele A and light blue are true counts
of allele B. Here, segments 2 and 4 are flipped. (D) Genome-specific copy number profiles c1 = [c1,1, c1,2, c1,3, c1,4] and c2 = [c2,1, c2,2, c2,3, c2,4] inferred under the assumption that the sample is heterogeneous, but the reference genome is haploid. Here, the entry ci,j for a segment j and genome Gi is the sum 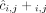 of allele-specific copy number values in a genome Gi. (E) Allele- and genome-specific copy number matrices
of allele-specific copy number values in a genome Gi. (E) Allele- and genome-specific copy number matrices 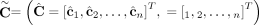 inferred under the assumption that the sample is heterogeneous and the reference genome is diploid. Segments 2 and 4 are
flipped alleles:
inferred under the assumption that the sample is heterogeneous and the reference genome is diploid. Segments 2 and 4 are
flipped alleles: 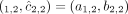 and
and 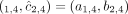 .
.











