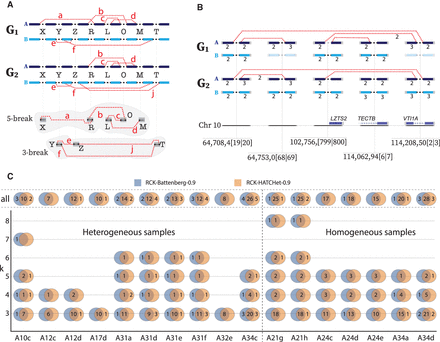
Evidence of complex k-break (k ≥ 3) rearrangements in metastatic prostate cancer. (A) Two complex rearrangements across two genomes in a heterogeneous sample. A 5-break rearrangement that produced four novel adjacencies {a, b, c, d} involving five reference adjacencies (X, R, L, O, and M), with novel adjacency a not present in genome G2. A 3-break rearrangement that produced three novel adjacencies {e, f, j} involving three reference adjacencies (Y, Z, and T), with novel adjacency j not present G1. (B, top) A complex 5-break rearrangement on Chromosome 10 in the karyotype inferred by RCK on sample A31a. Only the four novel adjacencies, five reference adjacencies, and incident segments involved in the rearrangement are shown. Copy numbers ≤1 are omitted for clarity, and absent segments/adjacencies are shown as faded. (Bottom) The locations of the corresponding double-stranded DNA breakages for the 5-break on Chromosome 10, indicated as x|y for each reference adjacency {(x)h, (y)t}. Three reference adjacencies lie in/near genes: reference adjacency 102,756,[799|800] falls within the promoter region for gene LZTS2; reference adjacency 114,208,50[2|3] falls inside gene VTI1A; and reference adjacency 114,062,94[6|7] falls inside gene TECTB. (C) Number of complex k-break (k ≥ 3) rearrangements reported in RCK-reconstructed karyotypes using HATCHet and Battenberg copy number inputs with novel adjacency utilization parameter P = 0.9. Values of 0 are omitted for clarity.











