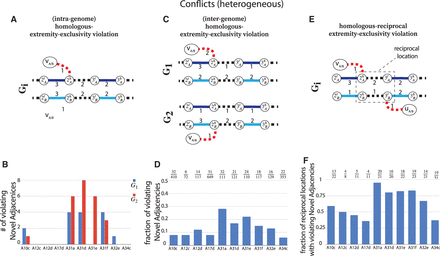
ReMixT karyotypes from heterogeneous prostate cancer samples have numerous violations of the generalized infinite sites constraints.
In A, C, and E, solid edges represent segment edges, black-dashed edges represent reference adjacency edges, and red dashed edges represent
novel adjacency edges. Integer values indicate copy numbers of corresponding segment and adjacency edges. (A) An intra-genome violation of the homologous-extremity-exclusivity constraint. To achieve copy number balance, both homologous
vertices 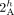 and
and 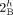 from genome Gi must be involved in novel adjacencies. (B) Number of novel adjacencies that violate the intra-genome homologous-extremity-exclusivity constraint in each cancer karyotype
inferred by ReMixT in each sample. (C) An inter-genome violation of the homologous-extremity-exclusivity constraint. To achieve copy number balance, both homologous
vertices
from genome Gi must be involved in novel adjacencies. (B) Number of novel adjacencies that violate the intra-genome homologous-extremity-exclusivity constraint in each cancer karyotype
inferred by ReMixT in each sample. (C) An inter-genome violation of the homologous-extremity-exclusivity constraint. To achieve copy number balance, both homologous
vertices 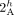 and
and 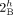 (in different genomes) must be involved in novel adjacencies. (D) The fraction x/y, where x is the number of novel adjacencies that violate the inter-genome homologous-extremity-exclusivity constraint (on at least
one of the extremities involved in a novel adjacency) in ReMixT karyotypes, and y is the total number of novel adjacencies reported by ReMixT as being present in both genomes. (E) A violation of the intra-genome homologous-reciprocal-extremity-exclusivity constraint. To achieve copy number balance,
both homologous-reciprocal vertices
(in different genomes) must be involved in novel adjacencies. (D) The fraction x/y, where x is the number of novel adjacencies that violate the inter-genome homologous-extremity-exclusivity constraint (on at least
one of the extremities involved in a novel adjacency) in ReMixT karyotypes, and y is the total number of novel adjacencies reported by ReMixT as being present in both genomes. (E) A violation of the intra-genome homologous-reciprocal-extremity-exclusivity constraint. To achieve copy number balance,
both homologous-reciprocal vertices 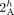 and
and 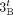 must be involved in novel adjacencies. Inter-genome violations of the homologous-reciprocal-extremity-exclusivity constraint
are also possible (Supplemental Fig. S17). (F) Fraction x/y, where x is the number of reciprocal locations with violations of either intra- or inter-genome (or both) homologous-reciprocal-extremity-exclusivity
constraint in ReMixT karyotypes; and y is the total number of reciprocal locations that both have novel adjacencies in ReMixT karyotypes.
must be involved in novel adjacencies. Inter-genome violations of the homologous-reciprocal-extremity-exclusivity constraint
are also possible (Supplemental Fig. S17). (F) Fraction x/y, where x is the number of reciprocal locations with violations of either intra- or inter-genome (or both) homologous-reciprocal-extremity-exclusivity
constraint in ReMixT karyotypes; and y is the total number of reciprocal locations that both have novel adjacencies in ReMixT karyotypes.











